Why we should not use Pandas Alone

Handling missing values is an important data preprocessing step in machine learning pipelines.
Pandas is versatile in terms of detecting and handling missing values. However, when it comes to model training and evaluation with cross validation, there is a better approach.
The imputer of scikit-learn along with pipelines provide a more practical way of handling missing values in cross validation process..
In this post, we will first do a few examples that show different ways to handle missing values with Pandas. After that, I will explain why we need a different approach to handle missing values in cross validation.
Finally, we will do an example using the missing value imputer and pipeline of scikit-learn.
Let’s start with Pandas. Here is a simple dataframe with a few missing values.
import numpy as np
import pandas as pd
df = pd.DataFrame(np.random.randint(10, size=(8,5)), columns=list('ABCDE'))
df.iloc[[1,4],[0,3]] = np.nan
df.iloc[[3,7],[1,2,4]] = np.nan
df

The isna function returns Boolean values indicating the cells with missing values. The isna().sum() gives us the number of missing values in each column.
df.isna().sum()
A 2
B 2
C 2
D 2
E 2
dtype: int64
The fillna function is used to handle missing values. It provides many options to fill in. Let’s use a different method for each column.
df['A'].fillna(df['A'].mean(), inplace=True) df['B'].fillna(df['B'].median(), inplace=True) df['C'].fillna(df['C'].mode()[0], inplace=True) df['D'].fillna(method='ffill', inplace=True) df['E'].fillna(method='bfill', inplace=True)
The missing values in columns A, B, and C are filled with mean, median, and mode of the column, respectively. For column D, we used ‘ffill’ method which uses the previous value in the column to fill a missing value. The ‘bfill’ does the opposite.
Here is the updated version of the dataframe:
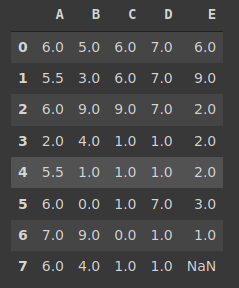
We still have one missing value in column D because we used the ‘bfill’ method for this column. With this method, the missing values are supposed to be filled with the values after them. Since the last value is a missing value, it was not changed.
The fillna function also accepts constant values. Let’s replace the last missing value with a constant.
df['E'].fillna(4, inplace=True)
As you have seen, the fillna function is pretty flexible. However, when it comes to train machine learning models, we need to be careful at handling the missing values.
Unless we use constant values, the missing values need to be handled after splitting the training and test sets. Otherwise, the model will be given information about the test set which causes data leakage.
Data leakage is a serious issue in machine learning. Machine learning models should not be given any information about the test set. The data points in the test sets need to be previously unseen.
If we use the mean of the entire data set to fill in missing values, we leak information about the test set to the model.
One solution is to handle missing values after train-test split. It is definitely an acceptable way. What if we want to do cross validation?
Cross validation means partitioning the data set into subsets (i.e. folds). Then, run many iterations with different combinations so that each example will be used in both training and testing.
Consider the case with 5-fold cross validation. The data set is divided into 5 subsets (i.e. folds). At each iteration, 4 folds are used in training and 1 fold is used in testing. After 5 iterations, each fold will be used in both training and testing.
We need a practical way to handle missing values in cross validation process in order to prevent data leakage.
One way is to create a Pipeline with scikit-learn. The pipeline accepts data preprocessing functions and can be used in the cross validation process.
Let’s create a new dataframe that fits a simple linear regression task.
df = pd.DataFrame(np.random.randint(10, size=(800,5)), columns=list('ABCDE'))
df['F'] = 2*df.A + 3*df.B - 1.8*df.C + 1.12*df.D - 0.5
df.iloc[np.random.randint(800, size=10),[0,3]] = np.nan
df.iloc[np.random.randint(800, size=10),[1,2,4]] = np.nan
df.head()

The columns A through E have 10 missing values. The column F is a linear combination of other columns with an additional bias.
Let’s import the required libraries for our task.
from sklearn.linear_model import LinearRegression from sklearn.pipeline import Pipeline from sklearn.impute import SimpleImputer
The SimpleImputer fills in missing values based on the given strategy. We can create a pipeline that contains a simple imputer object and a linear regression model.
imputer = SimpleImputer(strategy='mean')
regressor = Pipeline(
steps=[('imputer', imputer), ('regressor', LinearRegression())]
)
The “regression” pipeline contains a simple imputer that fills in the missing values with mean. The linear regression model does the prediction task.
We can now use this pipeline as estimator in cross validation.
X = df.drop('F', axis=1)
y = df['F']scores = cross_val_score(regressor, X, y, cv=4, scoring='r2')
scores.mean()
0.9873438657209939
The R-squared score is pretty high because this is a pre-designed data set.
The important point here is to handle missing values after splitting train and test sets. It can easily be done with pandas if we do a regular train-test split.
However, if you want to do cross validation, it will be tedious to use Pandas. The pipelines of scikit-learn library provide a more practical and easier way.
The scope of pipelines are quite broad. You can also add other preprocessing techniques in a pipeline such as a scaler for numerical values. Using pipelines allows automating certain tasks and thus optimizing processes.

